Image Processing
This part described how I manage to crush images into a handful of bit, and still have something recognizable.
Inspiration
I had in mind a demo I saw some times ago:
Evolution of Mona Lisa by Roger Johansson
It uses polygons to approximate the image. I wanted to use disk instead for aesthetic reasons. At the price of a less precise approximation.
Such algorithm already exists : Mona Lisa approximated with 150 circles. Although is this case, the author did not provide much implementation details.
Algorithm
The algorithm I used yield good results:


Which looks way better if you take a step back:


Let's take a look inside and figure out how it is working.
Genetic Algorithm you said?
Genetic programming is quite amazing. It's fun to explain because it relies on knowledge that everybody learn in hight school.
It's based on Darwin's evolution theory: Starting for a population of solution, let's mutate a little the solution at each generation and keep the best ones.
In order to use it we must:
Have a way to determine if a solution is better than another.
Describe the "gene" of a solution, a structure of data of quantified which can be altered.
Explicit how the solution gene mutate, to result in a slightly different solution.
data structure
To begin, we must define what is the "gene" of a solution: How do we describe the solution.
For our case, the gene will be an array of disks, each of theses disks have
a position
a radius
a color
Given a certain solution expressed with this data structure, we can draw the image solution.
And from there, we can compute the "fitness" which describe how good the solution is.
In our case, the fitness will be derived from the difference between the target image and the image solution.
Implementation
We have the big picture, now we let's dive into the implementation details.
Let's lay the rules:
1. How the fitness is computed
Here I compute the color distance for each pixel of the image. And sum the errors.
The distance is a simple euclidian distance in the rbg space.
We may argue that using a distance which respect the eye spec will be more relevant. ( the human eye tends to discern easily distance between bright colors than dark ones, or something like that )
I actually tried and did not had pertinent results with hsl space.2. How the gene mutate
Is the mutated gene close to the original ( soft mutation ), or is quite different ( hard mutation ) ?
This is a hard question, too close to the original and you may fall into local maximum, too far and you may never converge.
One technique derived from "simulated annealing" is to make hard mutation at start to explore the solution space, and as the generation goes, soften them to refine.

This illustration from wikipedia shows how simulated annealing avoid improving a local maximum. wikipedia simulated annealing
This is the solution implemented: after X mutation which does not improve the solution, use soft mutation instead.
The hard mutation either:
swap a disk with another one
random a disk ( set a random color, position and radius )
And the soft one either:
move the location to a close position
set the color, to a close one
alter the radius
3. How two genes are combined
In a classic genetic algorithm, one way to generate a solution at the N+1 generation is to take two solutions from the N generation and combined them together. However it did not makes much sense in our case and I decide to discard this mechanism.
As such, the only way to generate a new solution is by mutation.
4. How the gene pool is groomed
Should we keep only the better solutions ? How many "bad" solution should we keep ?
Once again this question is about "exploring the space vs refining a solution", with the danger of falling into local maximum or converge slowly.
In our case, the state of the art I found tends to keep only one solution in the gene pool.
I tries myself to use a larger genetic pool, without success.
I implemented a solution with a single element in the gene pool. At each generation, the mutation is either better and used, or worst and discarded. This simplify greatly the algorithm, although it's not really a genetic algorithm anymore, but something more like hill climbing algorithm.
Maybe I should have not give up on the genetic aspect this fast.
5. Incremental solution
Some algorithm I have found use a clever trick. They start with 8 disks, converge to a best solution, then add 8 another disk then converge, ect ...
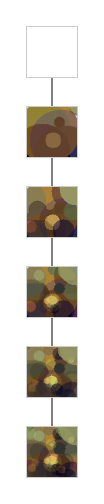

I had better results with this methods. And I got a way to monitor the progress.
Better, I used it to parallelize the algorithm :
starting from a blank solution, converge to a good solution with 8 disks
repeat to have N solution with 8 disks
take the best solution with 8 disk, to work a solution with 16 disk
repeat to have N solution with 16 disks
...
See ? now process can work isolated on making a solution converge. Sharing the result in a common solution tree.
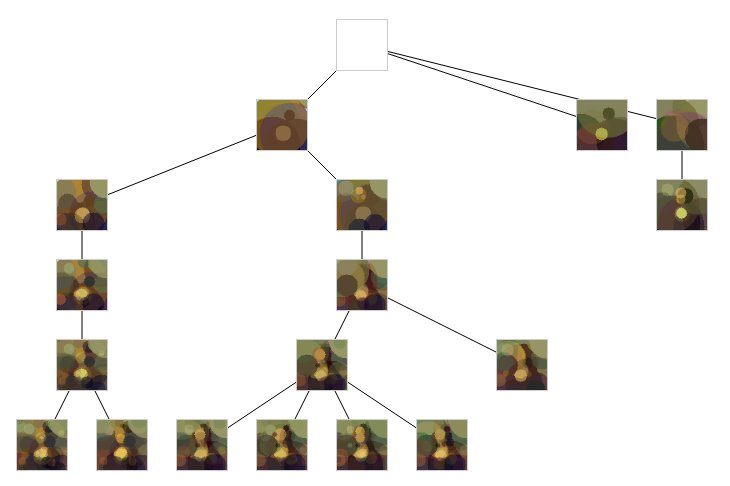
Cloudify
We have a solution which allows parallelization, let's spend some money on a big machine !
I use a redis cache to store the common solution, and spawn several machine in google compute engine to do the job.
while lamba calculus solutions ( aws lambda, gcp Function ... ) seems to fit here, they are not. The computation take some minutes, and will likely hit the timeout before end.
The flow is :
A process is launched
It get the solution tree, find a task to do ( take a N solution, make the N+8 solution converge )
make the solution converge
write the solution
repeat
And why not a front to read the solution tree and display the progress ?
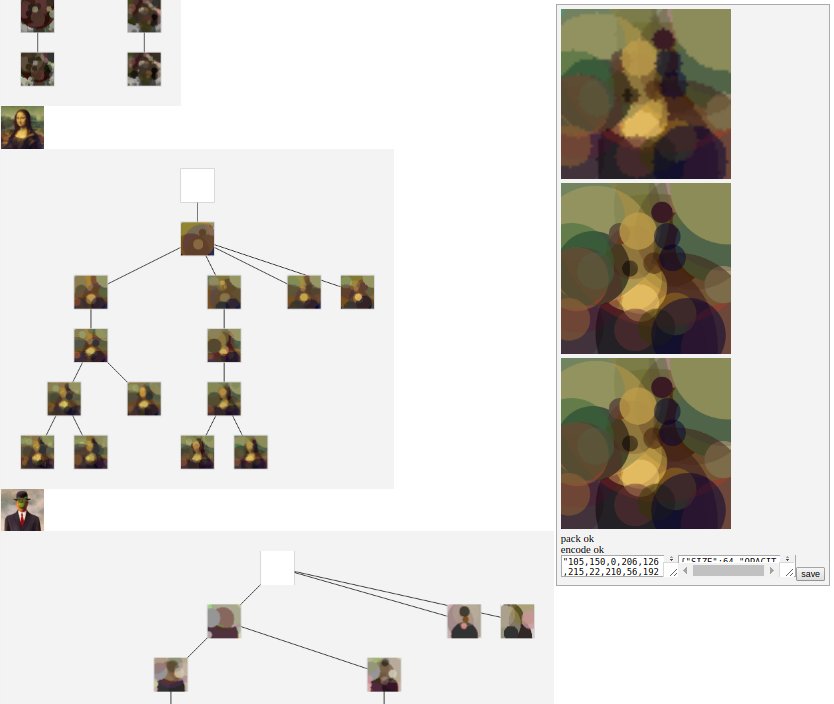
As you can see, we have several solutions as the tree leafs. I like to human pick the final solution, rather than take the one with the best fitness. To catch that little bit that the machine can't extract if you allows me to say.
Encoding
We are not done yet!
Remember the goal is to reduce the size of the asset to the minimum. Using JSON file is certainly not an option.
First, one thing I did not tell you is that the disk parameters are quantified:
the position is on a 64x64 grid
the color is contained in a palette of 256 elements
the color opacity is picked between 4 values
the radius is picket between 16 elements
So one disk can be encoded as bit array :
#### #### #### | #### #### | #### | ##
position color radius opacity
Which can be concatenated to represent the whole disk array.
Binary data in Js
Manipulating binary data is not so trivial in Js.
To do so, I write the binary as a file. It's possible with Uint8Array. With some tricks, I made the file downloadable from the app.
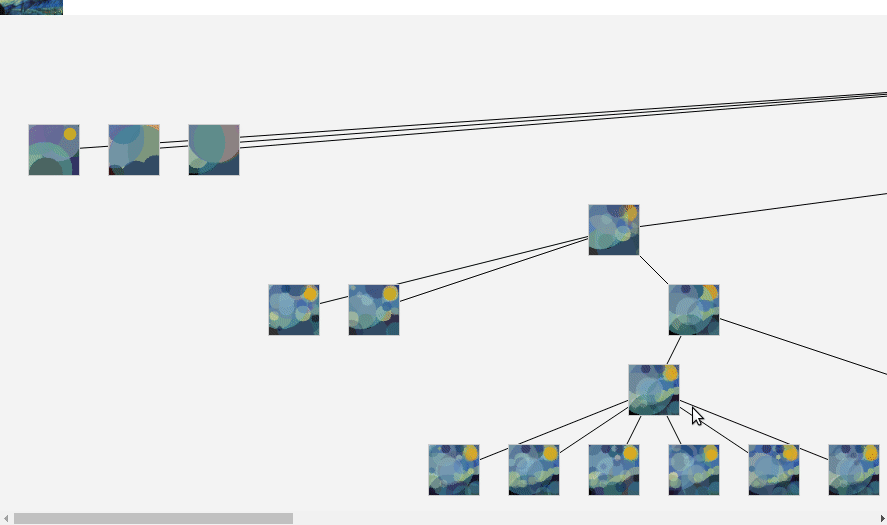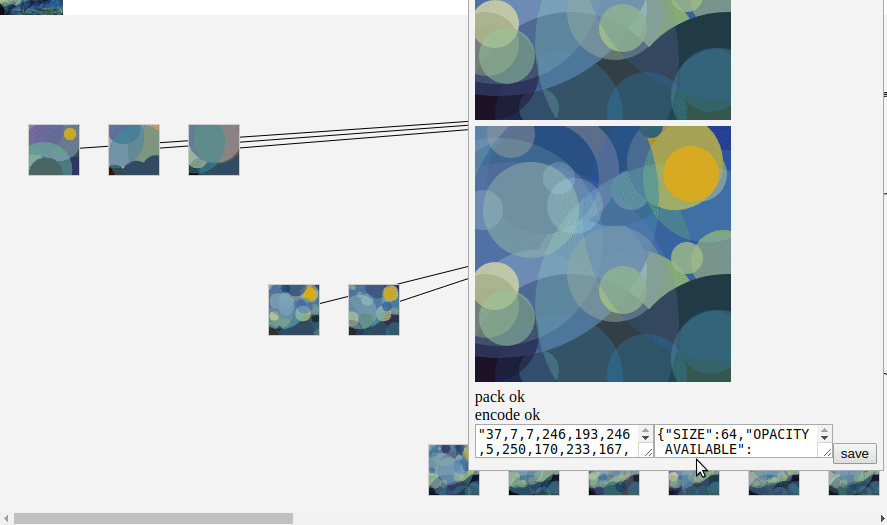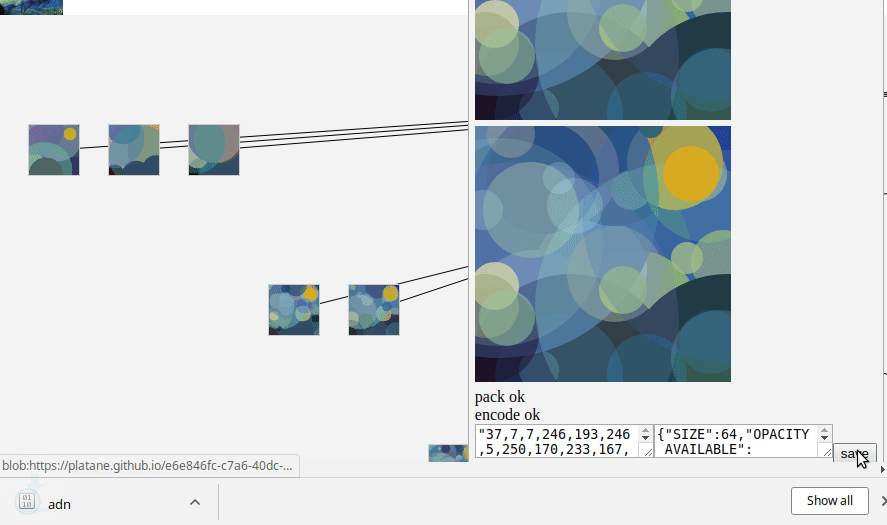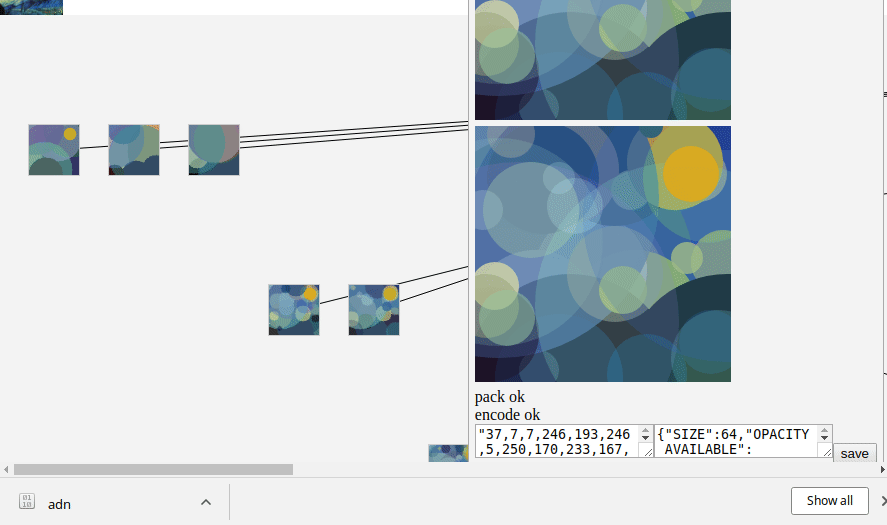
To read it, I use
fetch(fileUrl).then(res => res.arrayBuffer())
then it's a simple parsing to retrieve the original data.
